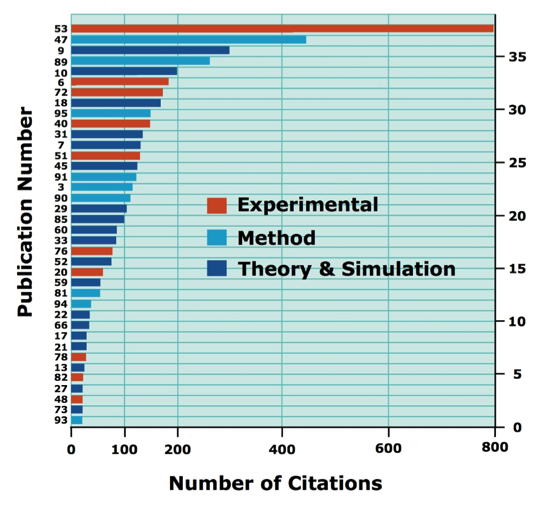
These statistics were originally compiled 09/2012
M primarily refers to an experimental tool or methodology.
T primarily refers to a theoretical observation based on structure or simulation.
798 | 53 | E | 1989 | Structural Origins of High Affinity Biotin Binding to Streptavidin |
444 | 47 | M | 1987 | Use of an Imaging Proportional Counter in Macromolecular Crystallography |
299 | 9 | T | 1976 | A Hypothetical Structure for an Intermolecular Electron Transfer Complex of Cytochromes c and b5. |
262 | 89 | M | 2001 | High Density Miniaturized Thermal Shift Assay as a General Strategy for Drug Discovery |
200 | 10 | T | 1977 | Structure and Function of Cytochrome c. |
184 | 6 | E | 1973 | The Structure of Oxidized Cytochrome c2 of Rhodospirillum rubrum. |
173 | 72 | E | 1992 | Crystallographic and Thermodynamic Comparison of Natural and Synthetic Ligands Bound to Streptavidin |
169 | 18 | T | 1980 | Structural and Functional Diversity in 4- a-Helical Proteins. |
150 | 95 | M | 2005 | Thermodynamic Stability Of Carbonic Anhydrase: Measurements Of Binding Affinity And Stoichiometry Using Thermofluor |
149 | 40 | E | 1985 | The Structure of Ferricytochrome c' from Rhodospirillum molischianum at 1.67 Å Resolution |
135 | 31 | T | 1983 | Structural Properties of Protein ß-Sheets |
131 | 7 | T | 1973 | Structural Bases for Function in Cytochrome c: An Interpretation of Comparative X-ray and Biochemical Data. |
130 | 51 | E | 1989 | Protein Crystal Growth in Microgravity |
125 | 45 | T | 1987 | Molecular Dynamics of a Cytochrome c-Cytochrome b5 Electron Transfer Complex |
123 | 91 | M | 2003 | Applications of Calorimetric Methods to Drug Discovery and the Study of Protein Interactions |
116 | 3 | M | 1972 | A Free Interface Diffusion Technique for the Crystallization of Proteins for X-ray Crystallography. |
112 | 90 | M | 2002 | Combinatorial Informatics in the Post-Genomics Era |
105 | 29 | T | 1982 | The a-Helix Dipole Model and Electrostatic Stabilization of a 4- a-Helical Proteins. |
100 | 85 | T | 1999 | Advances in diversity profiling and combinatorial series design. |
86 | 60 | T | 1990 | Competitor Analogs for Defined T Cell Antigens: Peptides Incorporating a Putative Binding Motif and Polyproline or Polyglycine Spacers |
85 | 33 | T | 1983 | Electrostatic Orientation During Electron Transfer Between Flavodoxin and Cytochrome c |
78 | 76 | E | 1994 | Structure-Based Design of Synthetic Azobenzene Ligands for Streptavidin |
76 | 52 | T | 1989 | Molecular Dynamics Simulation of a Phospholipid Micelle |
60 | 20 | E | 1981 | Crystallographic Structure of Rhodospirillum molischianum Ferricytochrome c' at 2.5 Å Resolution. |
55 | 59 | T | 1990 | A Molecular Dynamics Investigation of the Elastomeric Restoring Force in Elastin |
54 | 81 | M | 1997 | Serendipity meets precision: the integration of structure-based drug design and combinatorial chemistry for efficient drug discovery |
37 | 94 | M | 2005 | Decrypting the Biochemical Function of an Essential Gene from Streptococcus pneumoniae using Thermofluor Technology |
35 | 22 | T | 1981 | Conformational and Geometrical Properties of ß - Sheets in Proteins. III. Isotropically Stressed Configurations. |
34 | 66 | T | 1991 | The Identification of Tyrosine as a Common Key Residue in Unrelated H-2Kd Restricted Antigenic Peptides |
29 | 17 | T | 1979 | Conformations of Twisted Parallel ß-Sheets and the Origin of Chirality in Protein Structures. |
29 | 21 | T | 1981 | On the Evolutionary Relationship of 4- a-Helical Heme Proteins. |
28 | 78 | E | 1995 | Crystallographic and Thermodynamic Comparison of Structurally Diverse Molecules Binding to Streptavidin |
25 | 13 | T | 1977 | Structural Convergence During Protein Evolution |
23 | 82 | E | 1998 | In vitro evaluation and crystallographic analysis of a new class of selective, non-amide-based thrombin inhibitors. |
22 | 27 | T | 1982 | A Model for Catabolite Activator Protein Binding to Supercoiled DNA |
22 | 48 | E | 1988 | Molecular Factors Stabilizing Protein Crystals |
22 | 73 | T | 1992 | PROBIT: A Statistical Approach to Modeling Proteins from Partial Coordinate Data Using Substructure Libraries |
21 | 93 | M | 2003 | Chemical Genomics as an Emerging Paradigm for Postgenomic Drug Discovery |